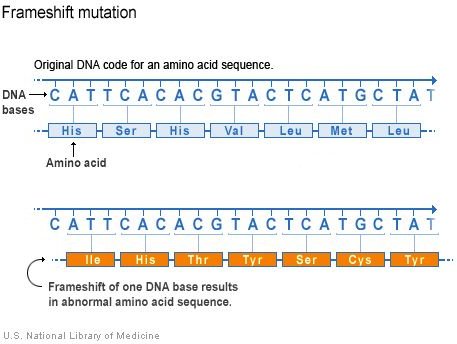
There are several classes of DNA mutations. Insertions and deletions are a type of mutation that changes the reading frame of a gene. This article discusses how they are introduced and how they can change affect a protein’s formula and function.
Opportunities for Mutations
DNA is copied by replication prior to cell division, or mitosis, and by transcription as the first step in gene expression. Opportunities for error arise any time that DNA is copied.
For example, if you were given a page of a book to copy, you may misspell or omit words. If you were writing in pencil, or typing on a keyboard, you could go back and correct your mistakes, greatly reducing the number of errors introduced during the process of copying. DNA polymerases have proofreading ability, similar to the backspace key on a keyboard. After proofreading, DNA polymerases make about one error per one million base pairs, which is impressive considering the workload.
Phrases like “She sells sea shells by the sea shore,” or “Toy boat,” said in rapid succession tie tonuges up in knots and put the speaker in a sputtering unintelligible mess. Similarly, when DNA polymerase encounters sections of repeating bases–such as a string of Gs or CAG repeats–it may literally fall off the DNA. If DNA polymerase does not pick up exactly where it left off, DNA may be added or omitted in the new DNA strand. These mutations are called insertions and deletions, respectively. If either occurs in a coding region, it will be a mutation that changes the reading frame of a gene.
Images
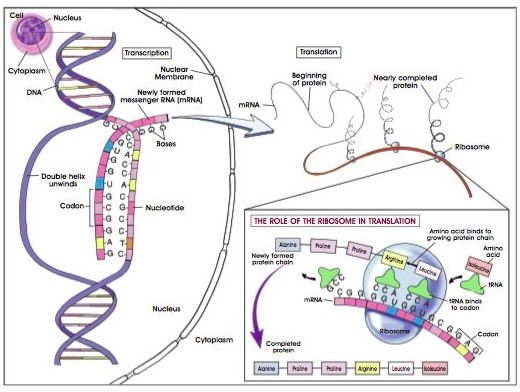
DNA to Protein
Inside the cell, proteins are the molecules that get things done. If the cell were a mafia ring, DNA would be the don and proteins would be the hitmen. The formula for each protein is built-in to the DNA sequence in groups called genes. Transcription is the first step in gene expression. During this process, one DNA strand (the template strand) is copied into RNA. Once DNA has been transcribed into messenger RNA (mRNA), the mRNA strand is processed, exported from the nucleus and translated into protein by the ribosome with the help of transfer RNA (tRNA).
The mRNA strand is read in groups of 3 nucleotides, called codons. A codon specifies which amino acid should be added to the peptide chain. Each amino acid has several codons that code for it; this can be a safeguard against mutations. Translation begins at the start codon, AUG (which codes for the amino acid methionine), a proceeds in a triplet pattern until one of three stop codons is encountered.
The newly formed protein is then subject to additional quality-control checks and post-translational modifications in the endoplasmic reticulum and golgi apparatus. The start codon sets the reading frame for translation. If insertion or deletion mutations are created in a multiple of three, translation will proceed “in-frame,” although one or several amino acids will be added or deleted depending on the type of mutation.
An insertion or deletion that does not occur in a multiple of three is a mutation that changes the reading frame of a gene. Translation will continue “out of frame” from the point of the mutation on. For example, an extra “a” added to “The cat ate the rat,” turns the sentence into “the caa tat eth era t,” if we keep the letters grouped in threes. Suddenly nothing makes sense because some words do not appear as they should.
The same thing happens on the peptide level: amino acids are incorporated into the peptide that should not be there. Frameshift mutations can create an early stop codon and result in a truncated protein, or eliminate a stop codon and the result in an abnormally long protein. Either way, the protein will mostly likely be non-functional.
Diseases Caused by Frameshift Mutations
-
Tay-Sachs disease is an autosomal recessive disorder marked by degeneration of both mental and physical capabilities. At its worst, death usually occurs around the age of four, although there are juvenile and late-onset forms of the disease. It is caused by the build-up of a particular sphingolipid in the brain, GM2 ganglioside, that is normally broken down by hexosaminidase. Though different mutations in the same gene can lead to the disease, in most cases, a 4 bp insertion in exon 11 of the HEXA gene (chromosome 15) renders a premature stop codon, leading to a profound deficiency of one of the subunits of hexosaminidase, hex A.
Advertisement -
Crohn’s disease is an inflammatory bowel disease that was previously thought to result from environmental factors. In 2001, Ogura et al. found a frameshift mutation in NOD2 (chromosome 16) that led to a cytosine insertion that ultimately reduced the protein’s activity.
It is important to note that not all frameshift mutations result in unstable, inactive proteins. Indeed, frameshift mutations can lead to new genes and novel proteins. After all, it is the ability to change DNA that leads to genetic adaptation and, ultimately, evolution of a species.
References
- Alberts B, Bray D, Johnson A, Lewis J, Raff M, Roberts K, Walter P. 1998. Essential Cell Biology: An Introduction to the Molecular Biology of the Cell. Garland Publishing, Inc: New York.
- Nussbaum RL, McInnes RR, Willard HF. 2001. Thompson & Thompson Genetics in Medicine, 6th ed. W.B. Saunders Company: Phildelphia.
- Ogura Y, Bonen DK, Inohara N, Nicolae DL, Chenn FF, Ramos R, Britton H, Moran T, Karaliuskas R, Duerr RH, Achkar JP, Brant SR, Bayless TM, Kirschner BS, Hanauer SB, Nuñez G, Cho JH. 2001. A frameshift mutation in NOD2 associated with susceptibility to Crohn’s disease. Nature 411: 603-606.
- “The process of gene expression” photo taken from https://stemcells.nih.gov/StaticResources/info/scireport/images/figurea6.jpg
- “Frameshift mutations alter a protein’s composition” taken from https://ghr.nlm.nih.gov/handbook/illustrations/frameshift.jpg