The combining of biochemical techniques with information technology has made it possible to understand higher levels of macromolecular structure. It also opens a vista of potential novel species.
Definition Plus Examples
Macromolecular modeling pinpoints atoms rather than parts of atoms, and features the study of large molecules such as proteins and nucleic acids by the manipulation of empirical data using computer software. It is done with the intent of gaining insight into higher levels of molecular structure or to study the interactions between molecules. Such study has sometimes been labeled “theoretical and computational biophysics.”
Some specific examples of macromolecular computer modeling (which covers all the behavior of macromolecules) are the thermodynamics of biological systems including protein folding, protein stability, conformational changes, enzyme catalysis and molecular recognition of DNA, RNA, proteins and membrane complexes. Modeling includes the interactions of atoms plus their interactions with van der Waals forces.
The so-called zero gradient calculation involves the minimizing of energy in such calculations. Such lower energy states are often involved in biological processes. Sometimes the presence of a solvent (such as water) is included in the calculations.
Examples of Computational Packages and Viewing Software
Some common varieties of computational software include Amber, Charmm, Gamess, MOPAC, Spartan, ConSurf and Sybyl. Viewers, often free, include PDB Viewer, PyMol, Chime, JMol and RasMol. Some of these forms are not only free, but are capable of being used on home computers. Some packages allow modeling of simple molecules without the input of empirically derived data. Others are best used by researchers who have gathered information to be entered as empirical data.
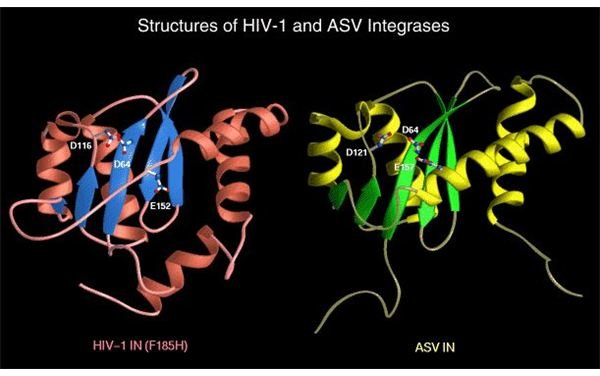
Modeled Macromolecular Structures
Credit: National Cancer Institute
Future Direction Prospects
One of the best prospects for future use of macromolecular computer modeling lies in the area of pharmacology. As an example, Merck scientists combined molecular modeling with crystallographic views to develop molecules that would serve as thrombin inhibitors. The multi-disciplinary combination of empirical study plus macromolecular computational modeling use seems to suggest that molecular modeling be used in the developing of new products to improve the human condition. Or, as the NSF reference (listed below) indicates, macromolecular modeling may be used to closely study molecular dynamics so that novel new substances can be synthesized to “develop safe, effective, precisely targeted medicines capable of relieving suffering and saving human lives.”
References and Resources
RCSB Protein Databank: A Resource for Studying Biological Macromolecules
Lycoming College: Macromolecular Modeling and Bioinformatics Page
Molecular Modeling and Simulation: An Interdisciplinary Guide