A ready reckoner of the best and most intuitive Open Source Academic Research Software. This list covers it all: graph plotting, bibliography, chemical structures, sequence analysis, algebra and mathematics - the entire gamut of academic research.
Academic scientific research has adapted the principles of ‘Open Source’ long before the term itself was coined. Good science has always been ‘open source’ - publications mention complete details of how the experiment was performed and any competent person with the right resources at his disposal would be able to reproduce the results. Hence the way to go in academic research has always been open source. Academic research software though, has traditionally been dominated by proprietary software. Most of the feature-rich software, especially those on the Windows platform, are not open source. But finding equally easy-to-use and powerful software which are open source is not too difficult. And for those scientists willing to embrace Open Source software for their research, this article simplifies their task all the more. Here’s a list of the Top Open Source Academic research software, covering a range of research needs. Go through them, download and install and try to use them in your work. You should be pleasantly surprised with the results!
1. Basic Essentials:
- Operating System: The most important component of open source academic research software has to be an open source operating system. And Linux is by far the most popular Open Source OS for the scientific community. It is the OS of choice for scientific applications from weather calculating super-computers to molecular dynamics and protein folding simulations. Red Hat Linux is the Enterprise distribution and Scientific Linux is a redistribution recompiled from the same source. Either of the two forms of Linux are used in a variety of physics, biophysics and computational biology laboratories the world over.
- Office Suite: After an Os the next essential for academic research is a good office suite. Most scientists use MS Word, Excel and Powerpoint to the extent that these have become almost generic terms for a word processor, spreadsheet and a presentation software. A very useful open source alternative is Open Office which provides all the required features of MS Office, at zero cost!
- Browser: With a lot of academic searches and data mining being done using online resources or databases, browsing the Net is an essential part of academic research. And yes, the default Internet Explorer that most scientists use is not open source. The most popular open source browser is Mozilla Firefox. Not only is it more secure and user friendly as compared to IE, one can add a plethora of ‘add-ons’ to customize it to our very specific needs. Google Chrome is partially open source, in the sense that the basic ‘Chromium’ its based on is open source, but additional APIs, the built-in PDF reader, etc. is not.
2. Bibliographic Tools:
-
JabRef is a reference management and bibliographic tool that acts as a front-end to BibTex. The interface is similar to its proprietary counterpart EndNote, making it quite easy to switch to. One can import references from online databases like PubMed and IEEEXplore, group references based on keywords, link to external files (documents, pdfs, etc., can import from or export to most common reference formats and also offers the cite-as-you-write feature. It is based on Java and hence is platform independent, capable of running on Windows, Mac or Linux.
Advertisement -
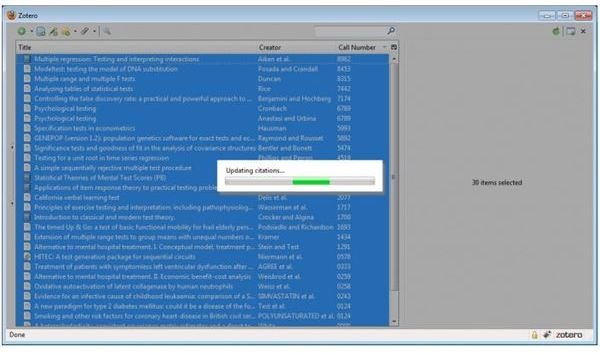
Zotero is browser-based tool, available as a plug-in for Mozilla Firefox, that can directly create your reference library as you search the web using your browser. One can not only store references, but also related files, website links, images, pdfs, etc. One can sync data with other computers, browse and access it from mobile devices like smartphones too. It easily integrates with Word or
OpenOffice, making it possible to simply drop in references on the fly. Multiple journal-specific styles are available freely.
Advertisement
3. Graph Plotting:
QtiPlot: An open source tools similar to Origin or SigmaPlot, QtiPlot can easily create publication quality 2D and 3D plots, including
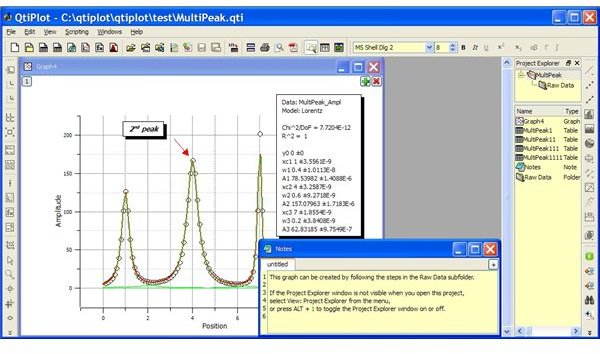
ding bar plots, pie charts, vector plots, contour and image plots with relevant error bars. Data is easily exported to multiple formats (EPS, PS, PDF, BMP, JPEG, TIFF, etc.). Linear and non-linear curve fitting of data, statistical analysis, data smoothing, convolution and deconvolution, matrices, correlation, integration and differentiation are its other features. Navigation is simple and powerful with data organized as project files allowing drag-and-drop and search facility.
4. Mathematics:
- Maxima is an algebra system, whose closest commercial equivalent would probably be Mathematica. It can be used for calculations involving integration, differentiation, matrices, differential equations, linear equations, polynomial expressions and 2D and 3D plots.
- A cross-platform numerical computational package, SciLab ) would prove to be an able open source competitor to MATLAB. It is particularly suitable for statistical analysis, image enhancement, simulating fluid dynamics, mathematical modeling, explicit and implicit dynamical system simulations (Xcos) and signal processing.
5. Physics:
Cernlib or the CERN Program Library, is a suite of data analysis tools, libraries and modules aimed at physicists involved in mathematical data modeling, detectors simulation, data handling and analysis. Most of these programs were written at CERN, the European Laboratory for Particle Physics.
6. Chemistry:
-
ChemDoodle : By far the closest in terms of features and ease of use to proprietary 2D/3D chemical structure drawing tools, ChemDoodle claims to do all that ChemDraw can do, and a little more, while retailing for a fraction of its price. It is available for Windows, Mac and Linux.
-
PyMol : Based on Python scripts, PyMol is one of the best molecular visualization tools, PyMol can also read crystallographic map files and provide a host of information to study molecules.
Advertisement -
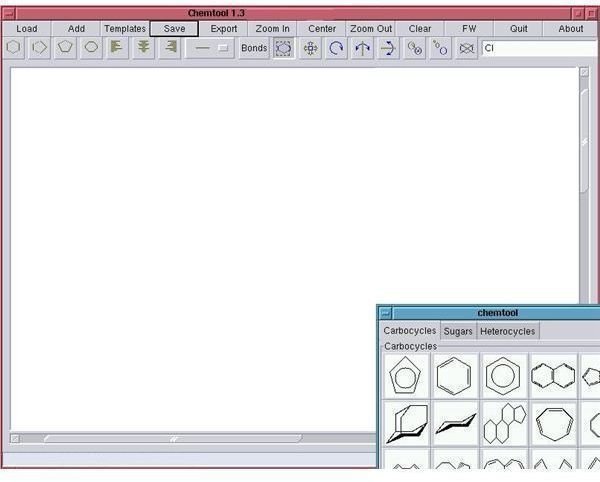
Chemtool : Chemtool is a small program useful for drawing chemical structures, on the Linux operating system. Though not as feature-rich as ChemDoodle, it still offers almost all features required for creating most 2D chemical structures.
 Advertisement
Advertisement
7. Biology:
EMBOSS is the European Molecular Biology Open Software Suite. This free suite is developed keeping in mind the needs of scientists in the field of molecular biology. One can easily perform protein and nucleic acid sequence alignments, database searching for patterns, motif and domain analysis and codon usage analysis, amongst other functions. User interface is intuitive and consistent across all the applications. It also easily integrates with other popular tools.
This list is definitely not exhaustive, as there are a lot of new and old open source academic research software tailored for every specific need. What this list does do is provide a starting point, to pick the best open source tool to get the job done, as compared to a proprietary software. i’d request all in the scientific community to at least try them out - there’s nothing to lose - and you may just realize that the thousands of dollars saved could be put for far more productive uses!