Typhoid fever is an illness caught by consuming food or drink contaminated with the bacteria Salmonella typhi. Finding out exactly which genes the organism needs to survive could speed up the development of new ways to tackle one of the major causes of disease in the developing world.
Typhoid Fever Symptoms
Polluted water is the main source of this bacterial infection and it’s a particular problem in the developing world. Typhoid fever is also a concern in South America and the Indian subcontinent.
A person with typhoid may display some of the following symptoms:-
- Severe headache
- Sudden onset of fever
- Profuse sweating
- Diarrhoea
- Abdominal pain
- Loss of appetite
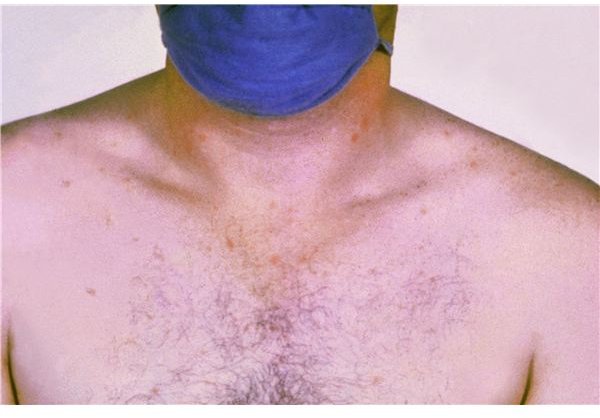
- Rash of rose-coloured spots
Treatment is usually with antibiotics and is nearly always successful. In most cases Typhoid fever is not fatal.
However, antibiotics are expensive and strains of Salmonella typhi are developing resistance to drug treatments. Therefore, there is an urgent need to understand more about the organism’s molecular biology to find new ways of attacking it.
Tackling the Cause of Typhoid Fever
An organism doesn’t necessarily need every single gene in its genome to survive. Some are essential, and others can be regarded as luxury items. Finding out which ones are essential can save a lot of research time by providing specific targets for new drug treatments.
Scientists from the Sanger Institute have developed a methodology where they can study every Salmonella typhi gene to work out which ones are essential for its survival and which ones are not. They discovered that 356 genes are for survival and 4162 are not.
To come up with this figure the researchers employed a technology called transposon-induced mutagenesis which they claim is much more powerful than any previous technique. A transposon is a mobile segment of DNA that can move to different positions in a genome. So what happens in this research is that a transposon will cut and paste itself into a genome and disable individual genes.
Genes for Survival
The scientists have called their new method TraDIS, which stands for Transposon Directed Insertion site Sequencing.
They inserted transposons into the S. typhi genome and then grew the bacteria to identify more than 300,000 insertion sites.
When one of these mobile genetic elements inserts into an essential gene it is silenced. The mutant cell doesn’t grow and so it and the transposon will be absent from the mutant pool. By sequencing the DNA from the entire gene pool, the researchers identify genes without transposons, and can therefore work out which one are essential for survival.
If a gene had taken up a transposon then it was not essential for survival. If a gene had no transposon within its DNA sequence then a functional copy of that gene is required for the bacteria to survive.
Gall Bladders
The technology was used to address a particular problem associated with S. Typhi. Namely that it can be spread by carrier humans who show no signs of the disease. The organism resides in human gall bladders which secretes bile. The bacteria can only survive here if they are tolerant to bile so the scientists went looking for the genes that conferred this tolerance.
They found 169 genes for bile tolerance, some of which were previously unknown. And so the researchers have highlighted many potential new targets for therapeutics and vaccines.